Discovering gene regulatory mechanisms underlying genomic imprinting and functions of imprinted, non-coding RNAs in neurophysiology and disease
“The Whipple laboratory at Harvard University utilizes molecular, cellular, and genomic approaches to determine how the genes inherited from our mother have a different effect on our brain than the genes inherited from our father.”
Genomic imprinting results in the preferential expression of a gene from either the maternal or paternal allele. Nearly all imprinted domains express non-coding RNAs, including long and small non-coding RNAs, in a parental biased manner. Yet the physiological functions and molecular mechanisms of these imprinted transcripts are poorly understood. With a focus on imprinted transcripts highly expressed in neurons, we seek to understand how parental gene inheritance influences gene expression in the brain through the activity of non-coding RNAs.
We have established an in vitro system that enables detection and manipulation of allele-specific expression in neurons. We are using this system to (1) delineate the neuronal networks regulated by imprinted microRNAs, (2) identify the neuronal targets of imprinted orphan small nucleolar RNAs, and (3) build a mechanistic understanding of cis regulation by imprinted long non-coding RNAs.
Imprinted disorders arise from inappropriate genetic alterations within imprinted domains. We have previously shown that targeting an imprinted long non-coding RNA is a viable therapeutic strategy for the treatment of an imprinted disorder, Angelman syndrome. Our laboratory is expanding on this work by examining the physiological role of imprinted non-coding RNAs during normal and dysregulated neurodevelopment.
Congratulations to Courtney on earning her PhD from the Harvard Program in Neuroscience! Over the years, she has grown into a rigorous scientist, aspiring bioinformatician, and thoughtful mentor. We can’t wait to see what she accomplishes next—look out world!
The Research Assistant will contribute to an individual research project within the lab’s broader program while also supporting laboratory operations. This position is ideal for candidates seeking significant hands-on research experience and mentorship in preparation for graduate training.
See full job description and submit application here https://us.smrtr.io/4KVbp
We are recruiting postdoctoral fellows to study the molecular functions of orphan snoRNAs, a large and poorly understood class of non-coding RNAs. Join us to uncover new mechanisms of RNA-mediated gene regulation in the brain. Read more about the positions here.
We’re excited that we have been awarded an R01 from NINDS to investigate the molecular consequences of snoRNA deletion in Prader-Willi syndrome.
Bongmin (Min) Bae, a postdoctoral researcher in the Whipple lab, has been awarded a new postdoctoral fellowship from the American Heart Association. The two-year award will support Bae’s research on how epigenetic mechanisms regulate gene expression in the mammalian brain.
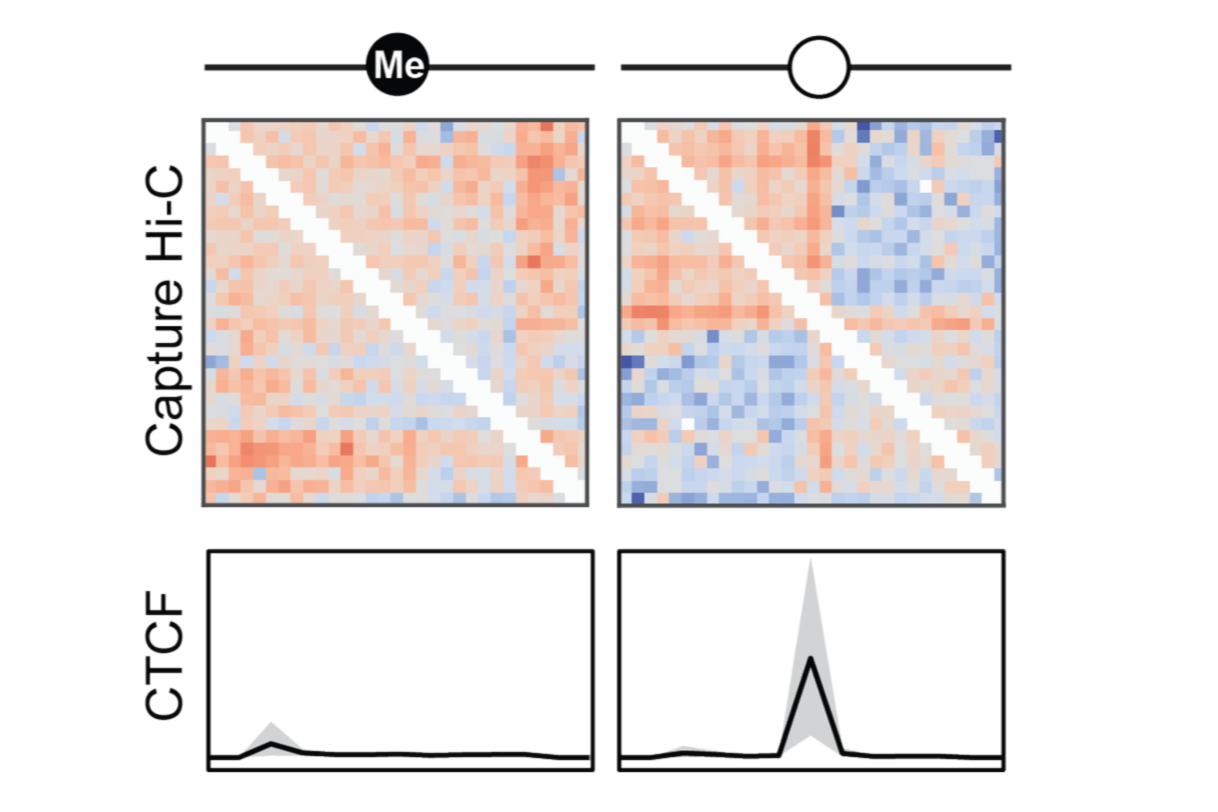
Our new preprint is now live on bioRxiv! We uncover how maternal and paternal genomes adopt distinct 3D chromatin structures across multiple imprinted domains in the mouse brain, and show that these architectures can work together with cis-acting long non-coding RNAs to shape allele-specific gene expression.
We invite all current G1 students in a HILS (Harvard Integrative Life Sciences) program to join us for an open house on November 4 at 6:15 PM. Dinner provided. Email for additional details.
Dina was selected as a finalist amongst a competitive pool of applicants and presented a five-minute lightning talk in front of 250+ participants in the Harvard research community.
Our proposal “Decoding Genomic Imprinting for Therapeutic Innovation,” developed together with lab member Dina Aljogol, has been selected for funding by the Milton Fund from Harvard University. We are grateful to the Harvard community for supporting this work.
Congratulations to two Whipple lab trainees, Courtney and Jen, for winning “the most beautiful experiment of the year” in the Harvard MCB Department! We also thank Min for her contributions to the original idea for the experiment; it was a truly collaborative effort. Courtney gave a talk at the departmental retreat highlighting this work.
Our latest study introduces a powerful chimeric eCLIP approach to map snoRNA-target RNA interactions across human and mouse cells. By analyzing binding partners of both core and accessory snoRNA-binding proteins, we uncover dozens of novel snoRNA interactions — shedding new light on their roles in RNA processing and modification. Read the full paper here.
Dina is a postdoctoral fellow interested in the molecular mechanisms of imprinted gene regulation. Before joining Harvard, she completed her PhD at Hamad Bin Khalifa University, where she studied chromatin organization and long-range promoter contacts in autism spectrum disorders.
The Whipple Lab welcomes MCO graduate student Isra Galicia Silva to the lab. She received her B.S. in Biochemistry and Microbiology from the Universidad del Valle de Guatemala and completed a Post-Baccalaureate at the University of Pennsylvania, where she worked with the J-Rad Lab on the interplay between metabolism, circadian rhythms, and cellular maturation.
Congratulations to Min Bae on receiving a postdoctoral fellowship award to further study how the Mest-Copg2 ICR achieves long-range regulation.
The Whipple Lab welcomes our newest member, Katherine Gu. Katherine joins us as a research assistant. She received her B.A. in Biochemistry and Chemical Biology from Vanderbilt University. Outside of lab, she enjoys working out, hiking, and trying restaurants in the Boston area.
Congratulations to Carlie McGrath Tydings who will be joining the Greenleaf Lab at the Stanford Department of Genetics in July, as a research assistant.
Congratulations to Aditya for graduating from Harvard! He will be attending graduate school at MIT in the fall.
Lily will be part of Harvard’s Program for Research Science and Engineering (PRISE) this summer. Her project with the Whipple lab will involve observing hallmarks of nucleolar stress and the role of small nucleoar RNAs on neuronal function.
Congratulations to Research Assistant Jen Yi! This fall, Jen will begin graduate school in the Harvard MCO program.
Debaleena Kar has joined the Whipple lab as a Postdoctoral Fellow. Her research focuses on the role of snoRNA in translation.
Daniel successfully defended his PhD thesis work on "Investigating the cis-regulatory mechanisms underlying neuronal imprinted expression." We thank Daniel for all his contributions to the lab and wish him the best of luck in his next career steps at Intellia Therapeutics!
This grant aims to investigate the role of SNORD116 in ribosome biology. For more details on this project and others funded by FPWR, a blog with the funded descriptions and a link to the recorded webinar presented by Dr. Theresa Strong, Director of Research Programs at FPWR can be found here.
A new study led by Daniel Loftus of the Whipple Lab has found that differences in maternal and paternal genomes in embryonic stem cells shape gene expression in neurons. The study, which appears in the latest issue of the journal Genes & Development, delves into the 3D structure of a genome region called Peg13-Kcnk9.
The Whipple lab enjoyed the MCB fall kick off celebration to welcome new students on campus. Graduate students interested in rotating should contact Amanda Whipple.
Congratulations to Courtney Whilden for being awarded an NIH F31 fellowship! This award will support her project on snoRNAs.
Carlie has been awarded HCRP (Harvard College Research Program) funding to pursue her work on the subnuclear localization of imprinted genes in the lab this summer.
Udbhav Chitta was the first member of the Whipple lab in 2019 and has been accepted into the biomedical graduate program at UMass Medical School. We are grateful to his contributions to the lab and are excited he will remain nearby. Warm wishes!
Whipple Lab Video Abstract
We aim to build mechanistic models of the cellular and physiological activity of imprinted non-coding RNAs, including microRNAs, small nucleolar RNAs, and long non-coding RNAs.
We are a collaborative group of scientists seeking to uncover new regulatory roles of RNA in the cell. Read our mission statement and learn more about who we are.
We are expanding our team! We aim to build a multidisciplinary group from diverse backgrounds, including computational biologists, molecular biologists, and neuroscientists.

































We’re excited to share that Christian Overbeck (Whipple Lab Intern 2019) has matched into the Internal Medicine/Global Health Residency Program at Stanford Health Care. Congratulations, Christian—we’re cheering you on!